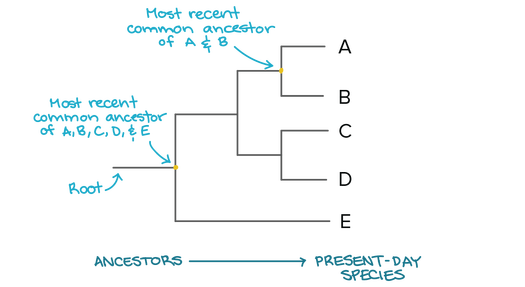
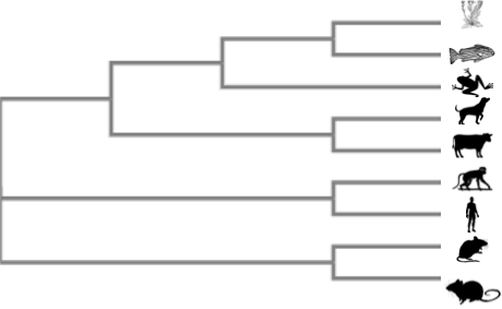
What is phylogeny?
Phylogenetics is the study of evolutionary history and relationships between organisms. To study these relationships, one can use a variety of inference methods which examine variable characters in the organisms being compared (Cavalli 1998). This can be used to determine the most recent common ancestors of various species as demonstrated above. Apart from species phylogeny, we can also use this technique to identify similarities and differences in protein sequences between species. Below are the steps used to develop a phylogenetic tree for the SHANK3 protein among various species.
Steps of Phylogenetic Tree Construction
(specific to SHANK3)
1) The most important step is the alignment of the DNA sequences in order to compare the data.
2) This data can then be constructed into a tree.
3) Next, the accuracy of the tree should be accessed. This can be done through bootstrap analysis in which a new multiple alignment is built by taking columns at random from the real alignment.
4) Finally, the a molecular clock can by used to assign dates to branches of the tree. This can be done by calculating the number of substitutions for a pair of homologous genes from a human and orangutan for example (Brown 2002).
Discussion
Analyzing protein phylogenetics provides us with yet another way of assessing the similarities and differences between the SHANK3 gene of species. When discussing SHANK3 protein homology, we compared the species' protein homologs. It is useful to note that the percent identities as indicated by homology differences do not necessarily correlate to the evolutionary history. For example, the percent identity comparing the human and mouse SHANK3 protein is higher than the percent identity comparing humans to the rhesus monkey. Evolutionarily however, the human SHANK3 protein is most closely related to the rhesus monkey.
It therefore may be informative to examine whether or not intelligence among species has a stronger correlation to protein homology or protein phylogenetics and what this may tell us about the mechanism in which the gene is regulated and expressed.
It therefore may be informative to examine whether or not intelligence among species has a stronger correlation to protein homology or protein phylogenetics and what this may tell us about the mechanism in which the gene is regulated and expressed.
References:
Brown TA. Genomes. 2nd edition. Oxford: Wiley-Liss; 2002. Chapter 16, Molecular Phylogenetics. Available from: https://www.ncbi.nlm.nih.gov/books/NBK21122/
Cavalli-Sforza LL. The DNA revolution in population genetics. Trends Genet. (1998);14:60–65.
Header:
https://rafaelbfalconi.deviantart.com/art/Tree-of-life-Concept-Art-Black-and-White-Version-377174915
Brown TA. Genomes. 2nd edition. Oxford: Wiley-Liss; 2002. Chapter 16, Molecular Phylogenetics. Available from: https://www.ncbi.nlm.nih.gov/books/NBK21122/
Cavalli-Sforza LL. The DNA revolution in population genetics. Trends Genet. (1998);14:60–65.
Header:
https://rafaelbfalconi.deviantart.com/art/Tree-of-life-Concept-Art-Black-and-White-Version-377174915
This web page was produced as an assignment for Genetics 564, an undergraduate capstone course at UW-Madison.