What is Protein Homology?
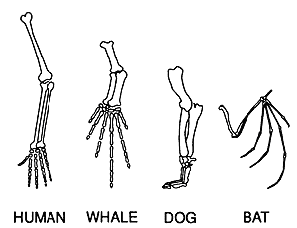
Homology is roughly defined as a common ancestry between two species as it relates to genes, proteins, or structures. Perhaps the most basic example of homology is structural homology. Structural homology is concerned with the structures of different species with a common ancestor or developmental origin. Homologous structures may differ in function between species. An example of this is the forelimbs of modern vertebrates (Fig 1). Gene homology refers to the common ancestry of a given DNA sequence. Events such as speciation or genetic duplication may cause differences in gene homology, referred to as homologs. The two types of homologs and orthologs and paralogs (Fig 2). Orthologs typically evolve as a result of speciation and generally retain their function. Paralogs, on the other hand, evolve as a result of duplication events and often vary in function (Lewis 2018). Finally, protein homology is often inferred from genetic sequence similarities between species. Protein homology can be determined using similarity searching programs including BLAST, FASTA, and SSEARCH. Found below are model organisms with proteins homologous to the human SHANK3 protein (Pearson 2013).
Homologous Structures
Fig 1. Forelimbs of Multiple Species
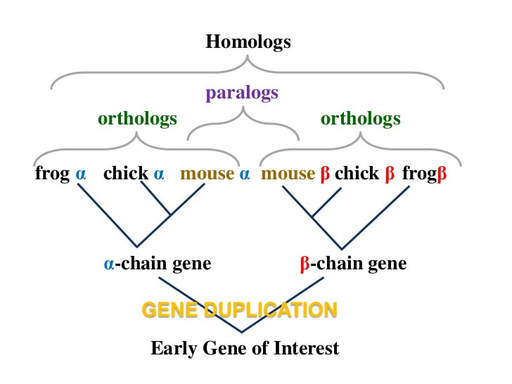
Homologous Genes
Fig 2. Difference between paralogs and orthologs indicated
Homologous Proteins
Homo sapiens (human) |
Mus musculus (mouse) |
Rattus norvegicus (rat) |
Macaca mulatta (rhesus monkey) |
Canis lupus familiaris (dog)
|
Bos taurus (cow)
|
Danio rerio (zebrafish) |
Xenopus tropicalis (frog) |
Arabidopsis thaliana (thale cress) |
Caenorhabditis elegans |
Drosophila melanogaster |
Discussion
Percent identity values as indicated above represent the degree of similarity between the SHANK3 protein sequence of the given model organism and homo sapiens. As percent identity increases, it would be expected that evolutionary distance would decrease (Stephen 2005). This seems to hold true with the SHANK3 homologs. The mammalian model organisms tend to have higher percent identities than the non-mammals. Because the SHANK3 protein is associated with intelligence, this presents an interesting opportunity for analysis. Homo sapiens (humans) are regarded as the most intelligent creatures in comparison to the other model organisms listed above. This poses the question of whether or not percent identities also correlate with intelligence levels of the listed species. The challenge in making this comparison is in measuring intelligence in non-human species. A test measuring g factor (see Home) would be ineffective in measuring the intelligence of a frog for example. Assuming that a standardized way to test for intelligence among all species could be created, the relationship between percent identity and intelligence may be of interest for future research.
References:
Chris Lewis (n.d.). Retrieved February 02, 2018, from http://homepage.usask.ca/~ctl271/857/def_homolog.shtml
Pearson, W. R. (2013). An Introduction to Sequence Similarity (“Homology”) Searching. Current Protocols in Bioinformatics / Editoral Board, Andreas D. Baxevanis ... [et Al.], 0 3, 10.1002/0471250953.bi0301s42. http://doi.org/10.1002/0471250953.bi0301s42
Stephen F. Altschul, John C. Wootton, E. Michael Gertz, Richa Agarwala, Aleksandr Morgulis, Alejandro A. Schäffer, and Yi-Kuo Yu (2005) "Protein database searches using compositionally adjusted substitution matrices", FEBS J. 272:5101-5109.
Stephen F. Altschul, Thomas L. Madden, Alejandro A. Schäffer, Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997), "Gapped BLAST and PSI-BLAST: a new generation of protein database search programs", Nucleic Acids Res. 25:3389-3402.
Header:
http://cdn.litlepups.net/2017/02/21/redirecting-to-articles-identification-chiens-chats-et-furets.jpg
Chris Lewis (n.d.). Retrieved February 02, 2018, from http://homepage.usask.ca/~ctl271/857/def_homolog.shtml
Pearson, W. R. (2013). An Introduction to Sequence Similarity (“Homology”) Searching. Current Protocols in Bioinformatics / Editoral Board, Andreas D. Baxevanis ... [et Al.], 0 3, 10.1002/0471250953.bi0301s42. http://doi.org/10.1002/0471250953.bi0301s42
Stephen F. Altschul, John C. Wootton, E. Michael Gertz, Richa Agarwala, Aleksandr Morgulis, Alejandro A. Schäffer, and Yi-Kuo Yu (2005) "Protein database searches using compositionally adjusted substitution matrices", FEBS J. 272:5101-5109.
Stephen F. Altschul, Thomas L. Madden, Alejandro A. Schäffer, Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997), "Gapped BLAST and PSI-BLAST: a new generation of protein database search programs", Nucleic Acids Res. 25:3389-3402.
Header:
http://cdn.litlepups.net/2017/02/21/redirecting-to-articles-identification-chiens-chats-et-furets.jpg
This web page was produced as an assignment for Genetics 564, an undergraduate capstone course at UW-Madison.