SHANK3 and Autism Spectrum Disorder
Conclusions
Introduction
Autism spectrum disorder prevalence has increased by 600 percent in the past 20 years which makes it an important area of research. The neurodevelopmental disorder which affects 1/68 children presents itself in a large spectrum of symptoms. It is correlated with the mutation and methylation of multiple genes, one of which being SHANK3.
SHANK3 is vital in neurodevelopment and acts specifically on the post-synaptic membrane. Specifically, it up-regulates the positive regulation of synaptic transmission which leads to an increase in post-synaptic density. Overall, SHANK3 is well conserved among species. There is some variation in the amount of domains in the rat, plant, and worm species. For example, the rat is notably missing one Ank-2 and PDZ domain. The zebrafish on the other hand has almost complete conservation.
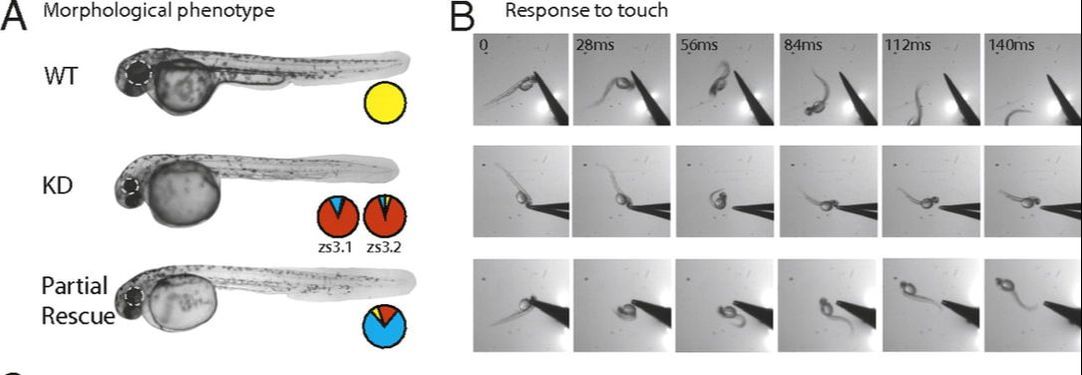
Apart from being well conserved, the zebrafish also has many traits which make it a great choice for a model organism. It is a vertebrate which has strong genetic similarity to humans. It is also transparent and develops quickly which increases the ease in which we are able to study the zebrafish. Most importantly however, the zebrafish has a clear mutant phenotype. Although there are some morphological differences after SHANK3 knockout, the more defined change is response to touch. Wild type zebrafish will swim away in response to touch, where mutant zebrafish will curl into a "ball" shape. After partial rescue, the zebrafish will instead swim toward the object that it was touched with (Figure 1). This makes SHANK3 disruption very easy to spot.
SHANK3 is vital in neurodevelopment and acts specifically on the post-synaptic membrane. Specifically, it up-regulates the positive regulation of synaptic transmission which leads to an increase in post-synaptic density. Overall, SHANK3 is well conserved among species. There is some variation in the amount of domains in the rat, plant, and worm species. For example, the rat is notably missing one Ank-2 and PDZ domain. The zebrafish on the other hand has almost complete conservation.
Apart from being well conserved, the zebrafish also has many traits which make it a great choice for a model organism. It is a vertebrate which has strong genetic similarity to humans. It is also transparent and develops quickly which increases the ease in which we are able to study the zebrafish. Most importantly however, the zebrafish has a clear mutant phenotype. Although there are some morphological differences after SHANK3 knockout, the more defined change is response to touch. Wild type zebrafish will swim away in response to touch, where mutant zebrafish will curl into a "ball" shape. After partial rescue, the zebrafish will instead swim toward the object that it was touched with (Figure 1). This makes SHANK3 disruption very easy to spot.
Figure 1. From top to bottom: wild type, knockout, and partial rescue morphology and response to touch
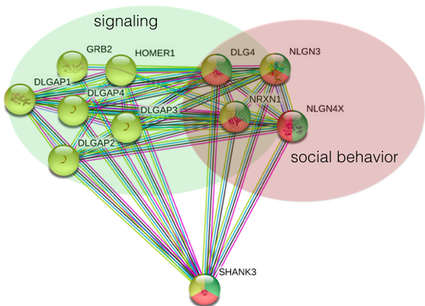
SHANK3's interaction network identifies a network which is highly involved in signaling and social behavior, both of which are impaired in autism patients (Figure 2).
Figure 2. Protein interaction network showing protein interactions important for signaling and social behavior
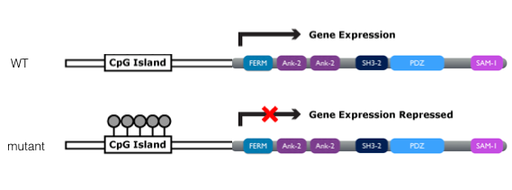
FiProtein interaction networks can be disrupted by methylation (Figure 3). Methylation is an epigenetic mechanism which often results in decrease of protein expression. SHANK3 methylation, as observed in autism spectrum disorder, results in a decrease in SHANK3 expression and decreases synapse function, specifically seen by a decrease in dendrite size and an abnormal response to touch. This occurs because DNA methylation recruits HDACs, or histone deacytltransferase complexes, which remove acetyl groups from histone tails, causing DNA to be more tightly wound around histones, thus lowering protein expression levels.
Figure 3. Affect of methylation of gene expression
My primary goal is to determine how methylation of SHANK3 affects synaptic function using zebrafish as a model. Zebrafish are excellent models for autism due to their genetic similarity to humans, transparency, quick development, and clear mutant phenotype, abnormal response to touch [4]. My hypothesis is that selectively targeting and knocking out methylation sites of SHANK3 will restore synapse function. My long term goal is to find a treatment for hyper-methylation of SHANK3 in autism patients in order to improve cognitive outcomes.
Aim 1: Determine conserved SHANK3 methylation sites necessary for synapse function
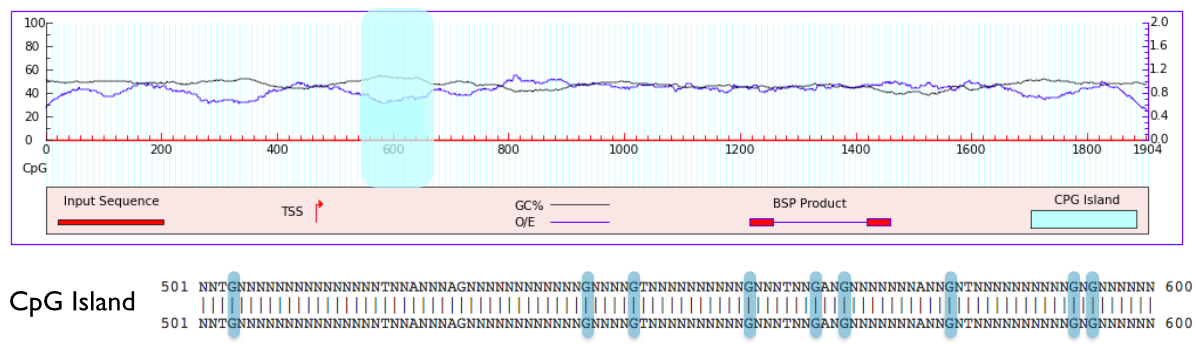
To determine conserved methylation sites, I first used MethPrimer to locate the CpG island in the promoter of SHANK3 and found this be somewhere between 500 and 600 base pairs. I then looked closely at this sequence and identified each potential CpG methylation site (Figure 4).
Figure 4. CpG island and potential CpG methylation sites shown in blue
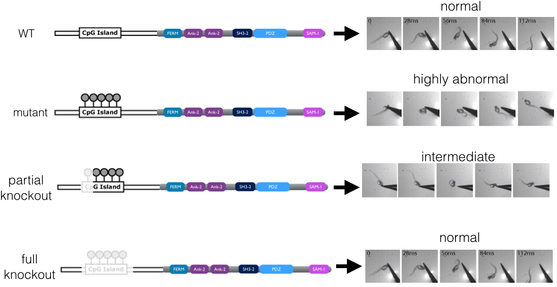
I will then knockout each CpG site as well as the entire CpG island and screen for abnormal response to touch. I hypothesize that knocking out methylation sites will result in restoration of normal response to touch. Therefore, I expect to see normal response to touch in wild type zebrafish, highly abnormal response to touch in mutant zebrafish, intermediate response to touch in the partial knockout zebrafish, and full restoration of normal response to touch in full knockout zebrafish (Figure 5).
Figure 5. Phenotypic outcome of wild type, mutant, partial knockout, and full knockout zebrafish
Aim 2: Determine how methylation affects synapse function in SHANK3 mutants
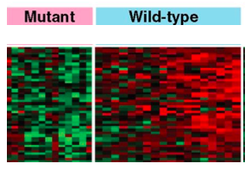
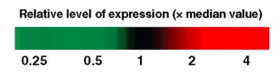
I will first perform RNA Seq on the mutant autism model and the wild type zebrafish in order to identify differentially expressed genes (Figure 6). When differentially expressed genes are identified, I will run these genes through gene ontology, looking specifically for signaling and social behavior. I will then use CRISPR/Cas9 to knockout each of these genes in zebrafish models and screen for abnormal response to touch. I hypothesize that SHANK3 methylation will result in differentially expressed genes involved in signaling and social behavior.
Figure 6. Relative level of expression in SHANK3 mutant (autism model) and wild type
Aim 3: Quantify SHANK3 interacting protein levels in wild type and mutant SHANK3 zebrafish
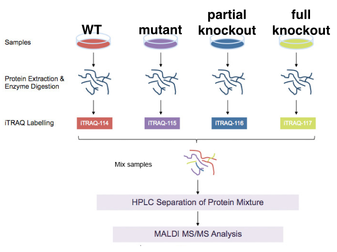
Using the wild type, mutant, partial knockout, and full knockout zebrafish from Aim 1, I will perform ITRAQ to quantify protein expression levels. Each group will be subject to protein extraction and enzyme digestion, ITRAQ labeling, mixture, separation, and mass spectrometry analysis (Figure 7).
Figure 7. ITRAQ protocol
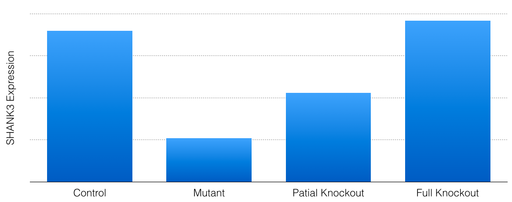
I will also once again screen for abnormal response to touch to confirm the expected phenotypes of each group. I hypothesize that mutant zebrafish will have higher levels of protein expression following SHANK3 methylation site knockout. This is because as methylation sites are knocked out, more HAT proteins will be recruited, and thus SHANK3 protein expression is expected to increase. I expect that the partial knockout zebrafish will experience only a partial restoration of expression levels, where full knockout of the CpG island will result in full restoration (Figure 8).
Figure 8. SHANK3 expression levels of control, mutant, partial knockout, and full knockout groups
Future Directions
After analysis of the results from this study, it would be helpful to determine the mechanism for the phenotypic outcomes of methylation. This will help us to explain how methylation correlates with the impaired signaling and social behavior seen in autism patients.
Additionally, if my hypotheses are correct, it would be reasonable to start testing the results of de-methylation in other species, and eventually in humans. De-methylation could prove to be an effective treatment in autistic humans and could alleviate some of the cognitive and social deficits that individuals experience.
Additionally, if my hypotheses are correct, it would be reasonable to start testing the results of de-methylation in other species, and eventually in humans. De-methylation could prove to be an effective treatment in autistic humans and could alleviate some of the cognitive and social deficits that individuals experience.
|
Draft One
|
Final Draft
|
|
| ||||||||||||
References
Dicke, U., & Roth, G. (2016). Neuronal factors determining high intelligence. Philosophical Transactions of the Royal Society B: Biological Sciences, 371(1685), 20150180. http://doi.org/10.1098/rstb.2015.0180
Li, S., Zhu, Y., Zhi, L., Han, X., Shen, J., Liu, Y., … Yang, X. (2016). DNA Methylation Variation Trends during the Embryonic Development of Chicken. PLoS ONE, 11(7), e0159230. http://doi.org/10.1371/journal.pone.0159230
SHANK3 gene - Genetics Home Reference. (2017, January). Retrieved January 31, 2018, from https://ghr.nlm.nih.gov/gene/SHANK3#sourcesforpage
Zhu, L., Wang, X., Li, X.-L., Towers, A., Cao, X., Wang, P., … Jiang, Y.-H. (2014). Epigenetic dysregulation of SHANK3 in brain tissues from individuals with autism spectrum disorders. Human Molecular Genetics, 23(6), 1563–1578. http://doi.org/10.1093/hmg/ddt547
F.F. Zimmermann, K.V. Gaspary, C.E. Leite, G. De Paula Cognato, C.D. BonanEmbryological exposure to valproic acid induces social interaction deficits in zebrafish (Danio rerio): a developmental behavior analysis. Neurotoxicol. Teratol., 52 (Part A) (2015), pp. 36-41
Yang, X., Hou, D., Jiang, W., & Zhang, C. (2014). Intercellular protein–protein interactions at synapses. Protein & Cell, 5(6), 420–444. http://doi.org/10.1007/s13238-014-0054-z
http://powerlisting.wikia.com/wiki/Intelligence_Manipulation
https://www.123rf.com/photo_18129529_autism-spelled-out-in-stacked-letter-blocks-with-copy-space-on-white-background.html
http://helicase.pbworks.com/w/page/17605615/DNA%20Methylation
http://philippe-plateaux.wix.com/illustrateur-medical#!medical
http://www.pnas.org/content/107/17/7863
http://science.sciencemag.org/content/352/6286/aaf2669https://www.savingcountrymusic.com/no-keith-urban-synth-and-drums-tracks-are-not-like-strings-and-chorus/human-being/
http://clincancerres.aacrjournals.org/content/20/12/3078
Header Image
https://www.parents.com/getting-pregnant/genetics/genetics-and-your-baby/
Dicke, U., & Roth, G. (2016). Neuronal factors determining high intelligence. Philosophical Transactions of the Royal Society B: Biological Sciences, 371(1685), 20150180. http://doi.org/10.1098/rstb.2015.0180
Li, S., Zhu, Y., Zhi, L., Han, X., Shen, J., Liu, Y., … Yang, X. (2016). DNA Methylation Variation Trends during the Embryonic Development of Chicken. PLoS ONE, 11(7), e0159230. http://doi.org/10.1371/journal.pone.0159230
SHANK3 gene - Genetics Home Reference. (2017, January). Retrieved January 31, 2018, from https://ghr.nlm.nih.gov/gene/SHANK3#sourcesforpage
Zhu, L., Wang, X., Li, X.-L., Towers, A., Cao, X., Wang, P., … Jiang, Y.-H. (2014). Epigenetic dysregulation of SHANK3 in brain tissues from individuals with autism spectrum disorders. Human Molecular Genetics, 23(6), 1563–1578. http://doi.org/10.1093/hmg/ddt547
F.F. Zimmermann, K.V. Gaspary, C.E. Leite, G. De Paula Cognato, C.D. BonanEmbryological exposure to valproic acid induces social interaction deficits in zebrafish (Danio rerio): a developmental behavior analysis. Neurotoxicol. Teratol., 52 (Part A) (2015), pp. 36-41
Yang, X., Hou, D., Jiang, W., & Zhang, C. (2014). Intercellular protein–protein interactions at synapses. Protein & Cell, 5(6), 420–444. http://doi.org/10.1007/s13238-014-0054-z
http://powerlisting.wikia.com/wiki/Intelligence_Manipulation
https://www.123rf.com/photo_18129529_autism-spelled-out-in-stacked-letter-blocks-with-copy-space-on-white-background.html
http://helicase.pbworks.com/w/page/17605615/DNA%20Methylation
http://philippe-plateaux.wix.com/illustrateur-medical#!medical
http://www.pnas.org/content/107/17/7863
http://science.sciencemag.org/content/352/6286/aaf2669https://www.savingcountrymusic.com/no-keith-urban-synth-and-drums-tracks-are-not-like-strings-and-chorus/human-being/
http://clincancerres.aacrjournals.org/content/20/12/3078
Header Image
https://www.parents.com/getting-pregnant/genetics/genetics-and-your-baby/
This web page was produced as an assignment for Genetics 564, an undergraduate capstone course at UW-Madison.